In my previous role at mProve Health, I read hundreds of protocols. As an electronic Clinical Outcome Assessment (eCOA) vendor, we were responsible for ensuring the right questionnaires were available at the right time and would be answered in a valid way. I didn’t appreciate how getting to a finished protocol is a delicate art in navigating research, regulatory precedence, and institutional knowledge. And complexity continues to grow: More regulations, more endpoints per trial, more amendments, and more options (in-person or remote assessments). The clinical protocol has become everything but the kitchen sink. So how can we simplify while maintaining data quality and integrity? We need reliable recommendations so that we can make decisions without wading through every piece of research and data available.
HumanFirst knows that ontologies are a great way to organize large datasets to build insights through computation. HumanFirst’s Atlas™ platform organizes >3k Digital Health Technologies (DHTs) across measures (passively-monitored COAs and digital biomarkers), medical conditions, and vendors. While DHTs are one piece of the puzzle, holistic protocol design requires more breadth. When selecting which COAs will represent study endpoints, a questionnaire, standardized task, DHT, or a combination thereof could be used to measure the same or related outcome:
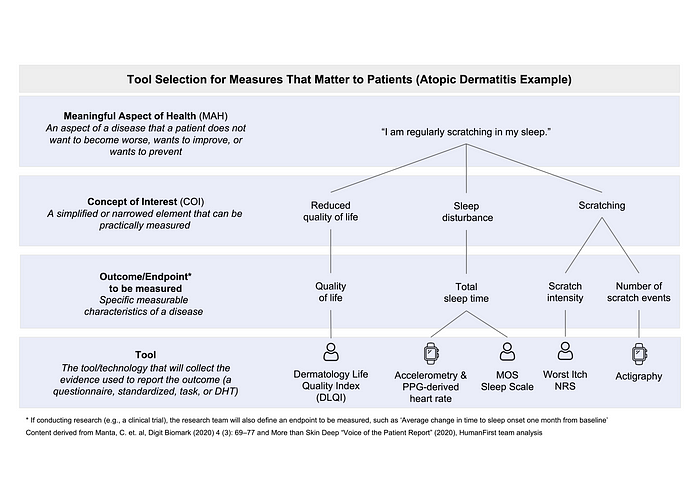
And yet, no single resource exists to compare them side-by-side to help you pick the right tool for the job. I started brainstorming with HumanFirst ways to expand the precision measurement capabilities of Atlas Network to include traditional, actively-collected COAs.
We knew that MAPI Research Trust’s ePROVIDE™ database maintains the world’s largest library of >5k actively-collected COAs. Turns out, five of the data elements in ePROVIDE™ and Atlas™ can be perfectly overlaid: Therapeutic area(s), medical condition(s), concepts of interest/domains, measures, and populations (Fig 2). These data elements create a backbone for a shared framework — or what we call “an aligned ontology”:
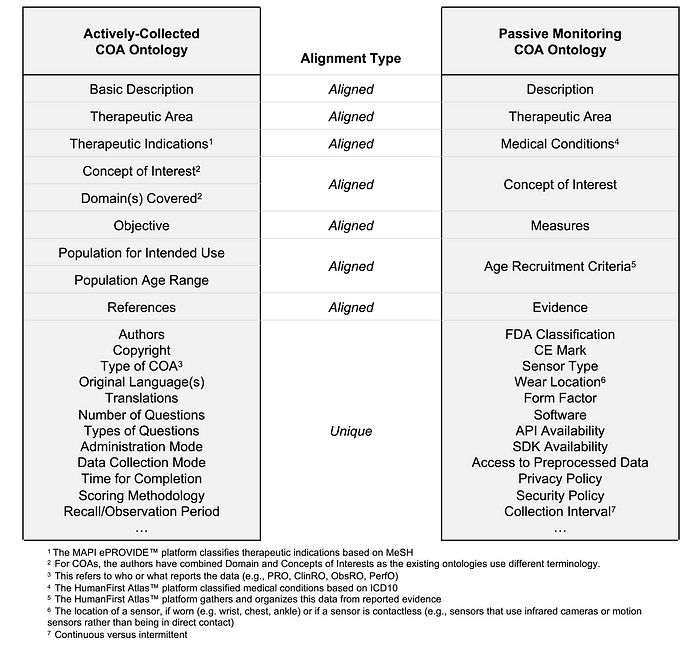
We validated this conceptual alignment and selected atopic dermatitis as a proof-of-concept. As described in our paper, this framework allows researchers to compare all relevant actively-collected and passively-monitored COAs and pick the best one for their study. Together with our co-authors from MAPI Research Trust, Syneos Health®, Roche Products Ltd, and Genentech Inc., we imagined how this aligned ontology could answer these questions:
- Which questionnaire(s), DHT(s), and standardized tasks have been used to measure COAs for atopic dermatitis?
- How has scratching behavior been measured actively and/or passively?
- Which tool(s) are well validated for my target population?
- Which tool(s) have data robust enough to support regulatory approval and reimbursement?
As well as practical considerations, such as:
- Can the tool be deployed (e.g., shipped, translated, operated, etc.) in the geographic region(s) needed for my trial?
As a fan of both actively-collected and passive-monitoring COAs, I don’t want or expect either to go away any time soon. But I do think this is a big step in helping the industry select the right tool for the job — such as picking a DHT to passively collect data when a PRO might be difficult to administer, or using both to contextualize passively-collected data to a participant’s reported experience.
This new way to research, identify, and select fit-for-purpose measures with confidence will reduce the number of endpoints in clinical studies without sacrificing quality and data integrity. Simpler protocols lessen the burden on participants and sites, can increase response rates and retention, and reduce issues with missing data along with many other downstream improvements to effort and cost.
Now imagine this aligned ontology persisted as a living repository mapping evidence collected as these decisions are made, the trials are conducted, and the results are published.
Additional datasets — like imaging, labs, or real-world data — can be added and aligned. This dataset could be used to train language models and/or generative AI, leading to insights and predictive recommendations for better drug discovery and development. This is one way our industry gets closer to the vision of precision measures for patient-centered drug development.
Special thanks to co-authors Christine Manta Campbell, Megan Parisi, Roya Sherafat-Kazemzadeh, Jessica Braid, Thomas Switzer, Marcelo Favaro, Caprice Sassano, Andy Coravos for their contributions and making this work possible.
Learn more about how HumanFirst is accelerating research using precision measures in their recent press release.
HumanFirst’s solutions have been used by leading academic medical institutions such as Harvard and Stanford, and by 24 out of 25 of the world’s largest pharmaceutical companies to unlock evidence-based precision measures that bring better treatments to market, faster.
Visit www.gohumanfirst.com to learn how you can Be First with HumanFirst.



.jpg)


